Research Themes
Exploring the boundaries of equilibrium
Our work spans non-equilibrium statistical mechanics, biological information processing, active matter, and the physics of learning and memory.
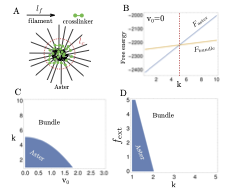
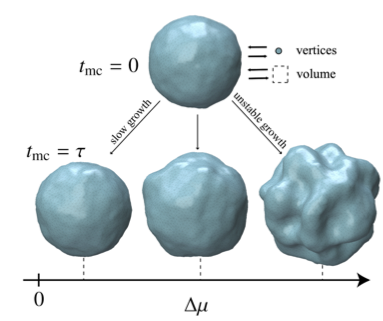
We use ideas from statistical mechanics, information theory, and machine learning to understand how physical and biological systems compute, adapt, and self-organize far from equilibrium.
Department of Chemistry · University of Chicago
Featured Research
Recent work from the group exploring how non-equilibrium dynamics gives rise to computation, learning, and functional structure.
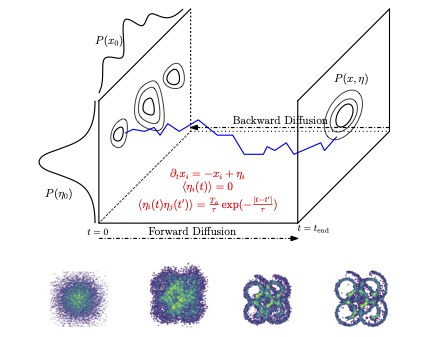
Driving generative diffusion out of equilibrium using active, temporally correlated noise creates a memory effect where high-level semantic information is stored in temporal correlations of auxiliary degrees of freedom.
What are the fundamental limits on the computations that non-equilibrium biophysical systems can perform? We establish rigorous bounds linking thermodynamic dissipation to computational power in chemical reaction networks and cellular processes.
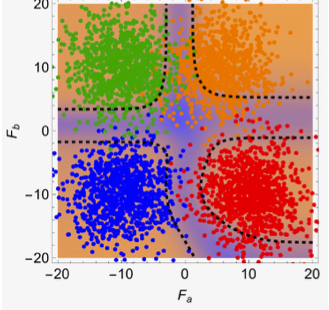
Combining high-throughput experiments with statistical mechanics theory, we map the diverse fitness landscape topologies governing protein-protein interactions, revealing how evolution navigates rugged energy landscapes.
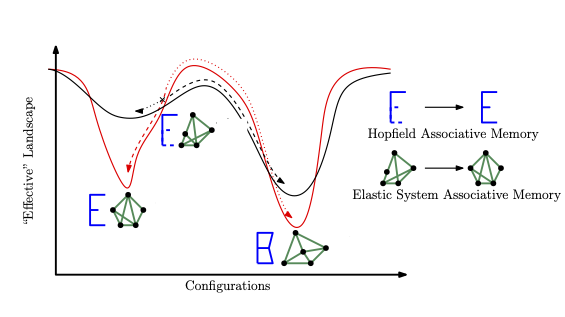
Drawing on the biology of neuromodulation, we design gated associative memory networks that dramatically extend memory retrieval capacity and exhibit emergent multistability—bridging neuroscience and the physics of learning.
We show that chemical reaction networks can perform in-context learning without attention mechanisms, bridging the gap between biochemistry and modern AI architectures.
Research Themes
Our work spans non-equilibrium statistical mechanics, biological information processing, active matter, and the physics of learning and memory.






Recent Highlights
A correspondence between Hebbian unlearning in neural networks and steady states generated by non-equilibrium dynamics.
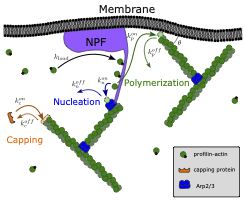
How cytoskeletal networks learn through mechanical feedback and active processes.
A review of the statistical mechanics underpinning memory, information storage, and retrieval in physical systems.